svyplan provides a simple and complete toolkit for
survey sample size determination. It covers sample sizes for proportions
and means, precision analysis, multistage cluster allocation,
multi-indicator optimization, strata boundary optimization, and power
analysis. All functions share a consistent interface built around
S3 generics, enabling round-trip conversions between sample
size and precision, and sensitivity analysis via
predict().
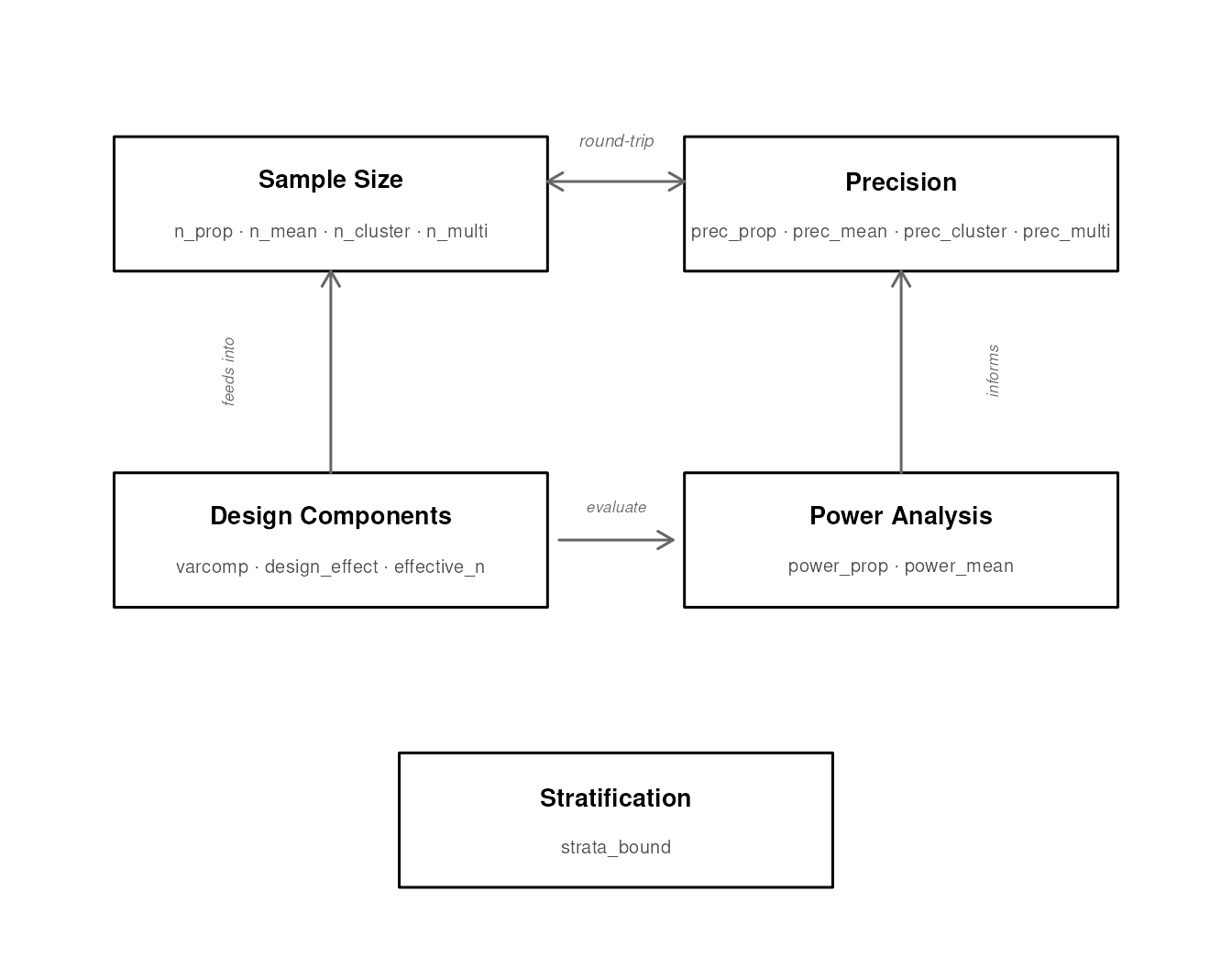
Function ecosystem
The package exports 18 functions organized into six families. The diagram below shows how they relate to each other.
| Family | Functions | Purpose |
|---|---|---|
| Plan | svyplan |
Reusable profile for design defaults |
| Sample size |
n_prop, n_mean,
n_cluster, n_multi, n_alloc
|
“How many units do I need?” |
| Precision |
prec_prop, prec_mean,
prec_cluster, prec_multi,
prec_alloc
|
“What precision does n achieve?” |
| Power |
power_prop, power_mean,
power_did
|
“Can I detect a change?” |
| Design |
varcomp, design_effect,
effective_n
|
Variance components and design effects |
| Strata | strata_bound |
Optimal stratification boundaries |
Every function returns an S3 object with print,
predict, and (where applicable) confint
methods.
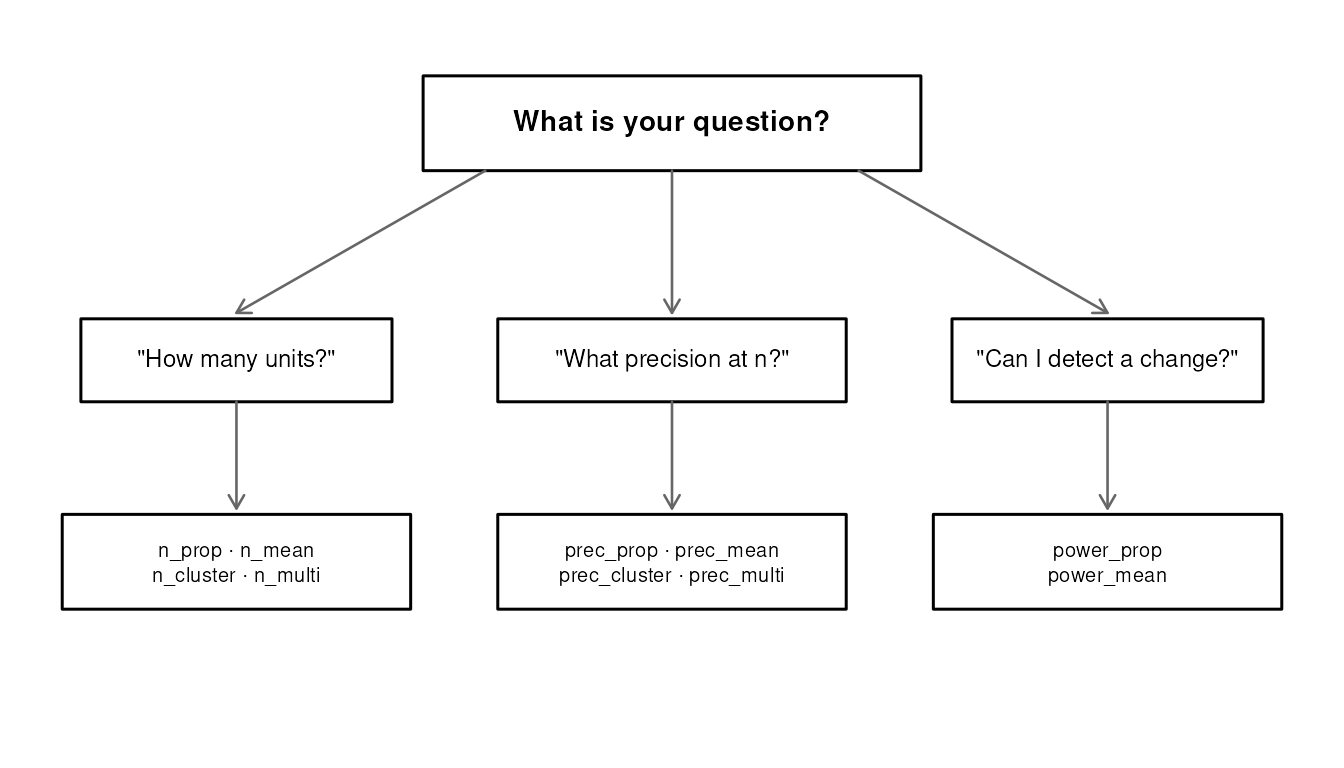
The n_*/prec_* duality
The most distinctive feature of svyplan is the bidirectional link
between sample size and precision functions. Every n_*
function has a prec_* counterpart, and they are exact
inverses.
Because n_* and prec_* are S3 generics, you
can pass a result object directly to its inverse. All design parameters
travel with the object:
# Compute sample size for a proportion
s <- n_prop(p = 0.3, moe = 0.05, deff = 1.5)
s
#> Sample size for proportion (wald)
#> n = 485 (p = 0.30, moe = 0.050, deff = 1.50)
# What precision does this n achieve?
p <- prec_prop(s)
p
#> Sampling precision for proportion (wald)
#> n = 485
#> se = 0.0255, moe = 0.0500, cv = 0.0850
# Recover the original sample size
n_prop(p)
#> Sample size for proportion (wald)
#> n = 485 (p = 0.30, moe = 0.050, deff = 1.50)You can also override a parameter on the way back:
n_prop(p, cv = 0.08)
#> Sample size for proportion (wald)
#> n = 547 (p = 0.30, cv = 0.080, deff = 1.50)Sample sizes for proportions and means
Proportions
Estimate vaccination coverage (expected ~70%) with a 5 percentage point margin of error:
n_prop(p = 0.70, moe = 0.05)
#> Sample size for proportion (wald)
#> n = 323 (p = 0.70, moe = 0.050)Three method options control how the margin of error is computed:
-
"wald"(default): the textbook normal approximation -
"wilson": Wilson score interval, better coverage near 0 or 1 -
"logodds": log-odds transformation, recommended for rare events
n_prop(p = 0.05, moe = 0.02, method = "wald")
#> Sample size for proportion (wald)
#> n = 457 (p = 0.05, moe = 0.020)
n_prop(p = 0.05, moe = 0.02, method = "wilson")
#> Sample size for proportion (wilson)
#> n = 469 (p = 0.05, moe = 0.020)
n_prop(p = 0.05, moe = 0.02, method = "logodds")
#> Sample size for proportion (logodds)
#> n = 475 (p = 0.05, moe = 0.020)Target a coefficient of variation instead of a margin of error:
n_prop(p = 0.70, cv = 0.05)
#> Sample size for proportion (wald)
#> n = 172 (p = 0.70, cv = 0.050)Means
Estimate average household expenditure (variance = 40,000, mean = 800) with a margin of error of 30:
n_mean(var = 40000, moe = 30)
#> Sample size for mean
#> n = 171 (var = 40000.00, moe = 30.000)Common parameters
Most single-stage n_*() and prec_*()
functions share these parameters:
| Parameter | Meaning |
|---|---|
deff |
Design effect multiplier (default 1) |
N |
Finite population size (enables FPC correction) |
alpha |
Significance level (default 0.05) |
resp_rate |
Expected response rate (inflates n by
1/resp_rate) |
n_prop(p = 0.70, moe = 0.05, deff = 1.5, N = 50000, resp_rate = 0.85)
#> Sample size for proportion (wald)
#> n = 566 (net: 481) (p = 0.70, moe = 0.050, deff = 1.50, resp_rate = 0.85)The design effect can be less than 1 for efficient designs
(e.g. well-stratified or PPS samples). svyplan accepts any
deff > 0.
n_cluster() / prec_cluster() use
cluster-specific inputs instead (stage_cost,
delta, rel_var, and optionally
resp_rate), and n_alloc() /
prec_alloc() have their own allocation-oriented
interfaces.
Survey plan profiles
When the same deff, N,
resp_rate, or alpha apply across many
single-stage calls, bundle them into a svyplan() profile to
avoid repetition:
plan <- svyplan(deff = 1.5, resp_rate = 0.85, N = 50000)
# Pass as a named argument
n_prop(p = 0.3, moe = 0.05, plan = plan)
#> Sample size for proportion (wald)
#> n = 566 (net: 481) (p = 0.30, moe = 0.050, deff = 1.50, resp_rate = 0.85)
n_mean(var = 100, moe = 2, plan = plan)
#> Sample size for mean
#> n = 170 (net: 144) (var = 100.00, moe = 2.000, deff = 1.50, resp_rate = 0.85)
power_prop(p1 = 0.30, p2 = 0.35, plan = plan)
#> Power analysis for proportions (solved for sample size)
#> n = 2328 (net: 1979, per group), power = 0.800, effect = 0.0500
#> (p1 = 0.300, p2 = 0.350, alpha = 0.05, deff = 1.50, resp_rate = 0.85)The profile can also hold cluster context (stage_cost,
delta) for n_cluster():
cl_plan <- svyplan(stage_cost = c(500, 50), delta = 0.05, resp_rate = 0.85)
n_cluster(cv = 0.05, plan = cl_plan)
#> Optimal 2-stage allocation
#> n_psu = 56 | psu_size = 14 -> total n = 784 (unrounded: 771.3894, net: 667)
#> cv = 0.0500, cost = 66551Piping works with both positional and named arguments:
plan |> n_prop(0.3, moe = 0.05)
#> Sample size for proportion (wald)
#> n = 566 (net: 481) (p = 0.30, moe = 0.050, deff = 1.50, resp_rate = 0.85)
plan |> n_prop(p = 0.3, moe = 0.05)
#> Sample size for proportion (wald)
#> n = 566 (net: 481) (p = 0.30, moe = 0.050, deff = 1.50, resp_rate = 0.85)
plan |> power_mean(5, var = 200)
#> Power analysis for means (solved for sample size)
#> n = 221 (net: 188, per group), power = 0.800, effect = 5.0000
#> (alpha = 0.05, deff = 1.50, resp_rate = 0.85)Explicit arguments always override plan defaults:
# plan has deff = 1.5, but deff = 2.0 wins here
n_prop(p = 0.3, moe = 0.05, plan = plan, deff = 2.0)
#> Sample size for proportion (wald)
#> n = 755 (net: 642) (p = 0.30, moe = 0.050, deff = 2.00, resp_rate = 0.85)Evaluating precision
The prec_*() functions answer the reverse question:
given a fixed sample size, what precision can you achieve?
prec_prop(p = 0.3, n = 400)
#> Sampling precision for proportion (wald)
#> n = 400
#> se = 0.0229, moe = 0.0449, cv = 0.0764
prec_mean(var = 40000, n = 300, mu = 800)
#> Sampling precision for mean
#> n = 300
#> se = 11.5470, moe = 22.6317, cv = 0.0144Design parameters work identically:
prec_prop(p = 0.3, n = 400, deff = 1.5, resp_rate = 0.85)
#> Sampling precision for proportion (wald)
#> n = 400 (net: 340)
#> se = 0.0304, moe = 0.0597, cv = 0.1015Confidence intervals
confint() extracts the expected confidence interval from
any sample size or precision object:
s <- n_prop(p = 0.3, moe = 0.05)
confint(s)
#> 2.5 % 97.5 %
#> 0.25 0.35
pr <- prec_prop(p = 0.3, n = 400)
confint(pr)
#> 2.5 % 97.5 %
#> 0.2550916 0.3449084
confint(pr, level = 0.99)
#> 0.5 % 99.5 %
#> 0.2409803 0.3590197For proportions, the interval is clamped to [0, 1].
Multi-indicator surveys
Household surveys like Demographic and Health Surveys (DHS) or
Multiple Indicator Cluster Survey (MICS) track many indicators
simultaneously. Each indicator has its own expected prevalence,
precision target, and design effect. n_multi() finds the
sample size that satisfies all targets at once.
targets <- data.frame(
name = c("stunting", "vaccination", "anemia"),
p = c(0.25, 0.70, 0.12),
moe = c(0.05, 0.05, 0.03),
deff = c(2.0, 1.5, 2.5)
)
n_multi(targets)
#> Multi-indicator sample size
#> n = 1127 (binding: anemia)
#> ---
#> name .n .cv_target .cv_achieved .binding
#> stunting 577 0.10204269 0.07297042
#> vaccination 485 0.03644382 0.02388518
#> anemia 1127 0.12755336 0.12755336 *The binding indicator (marked with *) drives the overall
sample size. The other indicators are estimated with better precision
than requested.
Achieved precision
prec_multi() computes the achieved precision for each
indicator at the chosen n:
plan <- n_multi(targets)
prec_multi(plan)
#> Multi-indicator sampling precision
#> name .se .moe .cv
#> stunting 0.02551067 0.05 0.10204269
#> vaccination 0.02551067 0.05 0.03644382
#> anemia 0.01530640 0.03 0.12755336Domain targets
Surveys often need adequate precision within subpopulations
(urban/rural, regions). Add domain columns to the targets data frame and
n_multi() optimizes each domain separately:
targets_dom <- data.frame(
name = rep(c("stunting", "vaccination", "anemia"), each = 2),
residence = rep(c("urban", "rural"), 3),
p = c(0.18, 0.32, 0.80, 0.60, 0.08, 0.16),
moe = c(0.05, 0.05, 0.05, 0.05, 0.03, 0.03),
deff = c(1.5, 2.5, 1.2, 1.8, 2.0, 3.0)
)
n_multi(targets_dom, domains = "residence")
#> Multi-indicator sample size (2 domains)
#> n = 1721 (binding: anemia)
#> ---
#> residence .n .binding
#> rural 1721 anemia
#> urban 629 anemiaDomain columns are specified explicitly via the domains
parameter. The returned n is the per-domain maximum (the
binding domain’s requirement), not the sum across domains. Use
$domains to see each domain’s sample size.
MICS/DHS-style relative margin of error
Programmes like UNICEF MICS and DHS express precision as a
relative margin of error (RME), defined as
MOE / p. This is related to the coefficient of variation by
RME = z * CV (at 95% confidence,
RME ≈ 1.96 * CV). To use svyplan with an RME target,
convert to an absolute margin of error: moe = RME * p.
# MICS-style: 12% RME per domain, domain-specific deff and prevalence
rme <- 0.12
targets_mics <- data.frame(
name = rep(c("stunting", "vaccination"), each = 3),
region = rep(c("North", "Central", "South"), 2),
p = c(0.35, 0.25, 0.40, 0.60, 0.75, 0.55),
deff = c(2.5, 2.0, 3.0, 1.5, 1.2, 1.8),
resp_rate = c(0.90, 0.85, 0.95, 0.90, 0.85, 0.95)
)
targets_mics$moe <- rme * targets_mics$p
n_multi(targets_mics, domains = "region")
#> Multi-indicator sample size (3 domains)
#> n = 1884 (binding: stunting)
#> ---
#> region .n .binding
#> Central 1884 stunting
#> North 1377 stunting
#> South 1264 stuntingThe MICS template reports sample size in households.
svyplan returns the number of individuals in the target
population. To convert, divide by the expected number of eligible
individuals per household:
n_hh = ceiling(n / (pb * hh_size)), where pb
is the share of the target population in the total population and
hh_size is the average household size.
Minimum sample size per domain
The min_n parameter sets a floor for the per-domain
sample size:
n_multi(targets_dom, domains = "residence", min_n = 300)
#> Multi-indicator sample size (2 domains, min_n = 300)
#> n = 1721 (binding: anemia)
#> ---
#> residence .n .binding
#> rural 1721 anemia
#> urban 629 anemiaMixing proportions and means
Targets can mix proportion and mean indicators. Use var
and mu columns for means (leave p as
NA), and p for proportions (leave
var/mu as NA):
targets_mixed <- data.frame(
name = c("vaccination", "expenditure"),
p = c(0.70, NA),
var = c(NA, 40000),
mu = c(NA, 800),
cv = c(0.05, 0.10),
deff = c(1.5, 2.0)
)
n_multi(targets_mixed)
#> Multi-indicator sample size
#> n = 258 (binding: vaccination)
#> ---
#> name .n .cv_target .cv_achieved .binding
#> vaccination 258 0.05 0.05000000 *
#> expenditure 13 0.10 0.02204793Multistage cluster designs
For two-stage or three-stage designs, the question is not just “how many?” but “how many clusters and how many units per cluster?” The answer depends on the cost structure and the degree of clustering.
Variance components from frame data
If frame data with cluster identifiers is available,
varcomp() estimates between-cluster and within-cluster
variance components:
set.seed(1)
n_clust <- 80
n_hh <- 15
cluster_id <- rep(seq_len(n_clust), each = n_hh)
cluster_mean <- rnorm(n_clust, mean = 50, sd = 10)
expenditure <- cluster_mean[cluster_id] + rnorm(n_clust * n_hh, sd = 20)
frame <- data.frame(cluster = cluster_id, expenditure = expenditure)
vc <- varcomp(expenditure ~ cluster, data = frame)
vc
#> Variance components (2-stage)
#> varb = 0.0398, varw = 0.1700
#> delta = 0.1896
#> k = 1.0589
#> Unit relvariance = 0.1981The delta value is the survey-planning measure of
homogeneity. It feeds directly into n_cluster() and the
cluster-planning mode of design_effect(). This
delta is not a generic mixed-model ICC:
varcomp() returns the bounded planning quantity from
Valliant, Dever, and Kreuter (2018), and that is the scale expected by
the cluster-planning functions. When delta is near 0, most
variation is within clusters, so the analytical cluster optimum tends to
favor many interviews in very few PSUs. When delta is near
1, most variation is between clusters, so the optimum tends to favor
very few interviews in many PSUs. Both are degenerate boundary cases for
the closed-form optimizer, so n_cluster() rejects values
numerically too close to 0 or 1.
Optimal two-stage allocation
Given per-stage costs and the variance structure,
n_cluster() finds the allocation that either minimizes cost
for a target CV, or minimizes CV for a given budget:
# Minimize cost to achieve CV = 0.05
n_cluster(stage_cost = c(500, 50), delta = vc, cv = 0.05)
#> Optimal 2-stage allocation
#> n_psu = 27 | psu_size = 7 -> total n = 189 (unrounded: 171.9801)
#> cv = 0.0500, cost = 21751
# Maximize precision within a budget
n_cluster(stage_cost = c(500, 50), delta = vc, budget = 100000)
#> Optimal 2-stage allocation
#> n_psu = 121 | psu_size = 7 -> total n = 847 (unrounded: 790.6791)
#> cv = 0.0233, cost = 100000Passing delta = vc (the varcomp object) automatically
extracts delta, rel_var, and
k.
Response rate in cluster designs
In budget mode, applying a response rate does not change the
allocation (the budget constrains the design); instead, the achieved CV
worsens. In CV mode, the first-stage sample is inflated by
1/resp_rate:
Fixed overhead costs
The standard cost model is purely linear:
C = c1*n_psu + c2*n_psu*psu_size. Real surveys also have a
fixed overhead (training, infrastructure, setup). The
fixed_cost parameter adds this term:
C = C0 + c1*n_psu + c2*n_psu*psu_size. In budget mode, only
budget - fixed_cost is available for the variable
component, reducing n_psu while leaving the optimal psu_size/n_psu ratio
unchanged. In CV mode, all sample sizes stay the same and
fixed_cost is simply added to the total cost:
# Budget mode: fixed overhead reduces the allocatable budget
n_cluster(stage_cost = c(500, 50), delta = vc, budget = 100000, fixed_cost = 5000)
#> Optimal 2-stage allocation
#> n_psu = 115 | psu_size = 7 -> total n = 805 (unrounded: 751.1452)
#> cv = 0.0239, cost = 100000 (fixed: 5000)
# CV mode: same allocation, cost increases by fixed_cost
n_cluster(stage_cost = c(500, 50), delta = vc, cv = 0.05, fixed_cost = 5000)
#> Optimal 2-stage allocation
#> n_psu = 27 | psu_size = 7 -> total n = 189 (unrounded: 171.9801)
#> cv = 0.0500, cost = 26751 (fixed: 5000)The same parameter is available in n_multi() for
multistage multi-indicator designs.
Evaluating cluster allocations
prec_cluster() computes the achieved CV for any
allocation:
alloc <- n_cluster(stage_cost = c(500, 50), delta = vc, budget = 100000)
prec_cluster(alloc)
#> Sampling precision for 2-stage cluster
#> n_psu = 121 | psu_size = 7 -> total n = 847
#> cv = 0.0233
# Arbitrary allocation
prec_cluster(n = c(50, 12), delta = 0.05)
#> Sampling precision for 2-stage cluster
#> n_psu = 50 | psu_size = 12 -> total n = 600
#> cv = 0.0508Design effects
The cluster design effect formula
deff = 1 + (psu_size - 1) * delta links cluster size to
precision loss:
design_effect(delta = 0.05, psu_size = 20, method = "cluster")
#> [1] 1.95After data collection, compute the realized design effect from survey weights:
set.seed(123)
w <- runif(500, 0.5, 4)
design_effect(w, method = "kish")
#> [1] 1.198255
effective_n(w, method = "kish")
#> [1] 417.2735Multi-indicator multistage
For surveys with multiple indicators in a cluster design, provide
delta_psu columns and a cost/budget to
n_multi():
targets_ms <- data.frame(
name = c("stunting", "vaccination"),
p = c(0.25, 0.70),
cv = c(0.10, 0.08),
delta_psu = c(0.05, 0.02)
)
n_multi(targets_ms, stage_cost = c(500, 50), budget = 80000)
#> Multi-indicator optimal allocation (2-stage)
#> n_psu = 68 | psu_size = 14 -> total n = 952 (unrounded: 927.2796)
#> cv = 0.0728, cost = 80000 (binding: stunting)
#> ---
#> name .n .cv_target .cv_achieved .binding
#> stunting 491.76043 0.10 0.0728 *
#> vaccination 84.08575 0.08 0.0241Optimal strata boundaries
When a continuous auxiliary variable is available on the frame,
strata_bound() finds the cut points that minimize the
sample size (or CV) for a given number of strata.
set.seed(12345)
x <- rlnorm(5000, meanlog = 6, sdlog = 1.2)
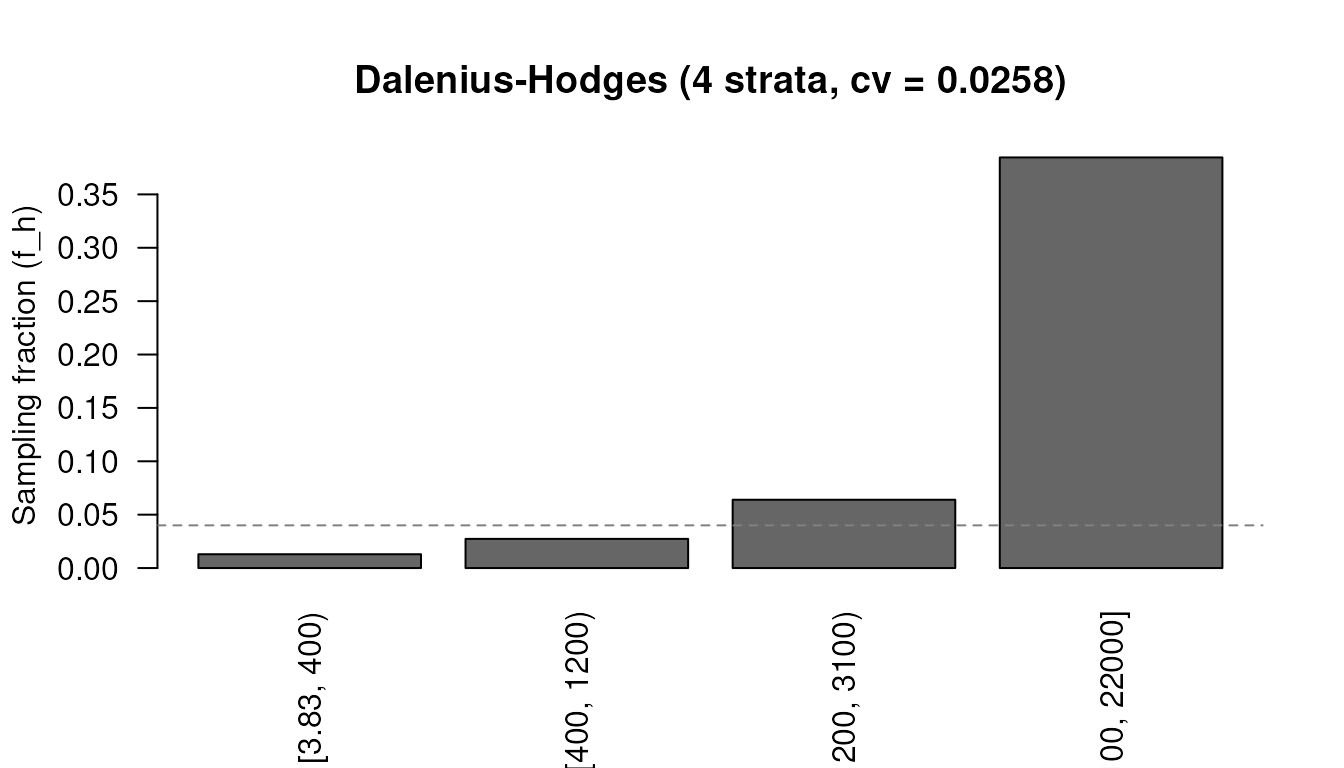
strata_bound(x, n_strata = 4, n = 300, method = "cumrootf")
#> Strata boundaries (Dalenius-Hodges, 4 strata)
#> Boundaries: 400.0, 1200.0, 3100.0
#> n = 300, cv = 0.0199
#> Allocation: neyman
#> ---
#> stratum lower upper N share sd n
#> 1 3.826706 400.00 2473 0.495 105.3 48
#> 2 400.000000 1200.00 1647 0.329 222.7 68
#> 3 1200.000000 3100.00 672 0.134 519.5 65
#> 4 3100.000000 21957.85 208 0.042 3094.3 119Four methods are available:
| Method | Algorithm | Speed | Best for |
|---|---|---|---|
"cumrootf" |
Dalenius-Hodges cumulative root frequency | Fast | General use |
"geo" |
Geometric progression | Fast | Skewed data |
"lh" |
Lavall0e9e-Hidiroglou iterative | Moderate | Skewed data |
"kozak" |
Kozak random search | Slow | Global optimum |
The Kozak method often finds better boundaries for highly skewed distributions:
strata_bound(x, n_strata = 4, cv = 0.05, method = "kozak")
#> Strata boundaries (Kozak, 4 strata)
#> Boundaries: 405.6, 1219.3, 3447.4
#> n = 59, cv = 0.0500
#> Allocation: neyman
#> ---
#> stratum lower upper N share sd n
#> 1 3.826706 405.6271 2494 0.499 106.6 10
#> 2 405.627119 1219.2788 1645 0.329 226.7 14
#> 3 1219.278753 3447.3536 690 0.138 589.2 15
#> 4 3447.353550 21957.8496 171 0.034 3206.8 20Allocation methods
Allocation within strata is controlled by the alloc
parameter:
-
"proportional": -
"neyman"(default): (minimizes national CV) -
"optimal": (accounts for costs) -
"power": Bankier (1988) compromise, , where controls the trade-off between national precision (, Neyman) and equal subnational CVs ()
# Bankier power allocation (compromise, power_q = 0.5)
strata_bound(x, n_strata = 4, n = 300, alloc = "power", power_q = 0.5)
#> Strata boundaries (Lavallee-Hidiroglou, 4 strata)
#> Boundaries: 456.3, 1354.4, 4360.4
#> n = 300, cv = 0.0216
#> Allocation: power (power_q = 0.50)
#> Converged: yes
#> ---
#> stratum lower upper N share sd n
#> 1 3.826706 456.3167 2710 0.542 120.7 34
#> 2 456.316687 1354.3749 1529 0.306 244.5 52
#> 3 1354.374865 4360.3970 650 0.130 743.3 103
#> 4 4360.396977 21957.8496 111 0.022 3441.5 111Classifying new observations
predict() applies the learned boundaries to new data,
returning a factor:
sb <- strata_bound(x, n_strata = 4, n = 200, method = "cumrootf")
new_x <- c(100, 500, 1500, 5000)
predict(sb, newdata = new_x, labels = paste0("S", 1:4))
#> [1] S1 S2 S3 S4
#> Levels: S1 S2 S3 S4Stratified allocation
Given a sampling frame with stratum population sizes (N)
and stratum standard deviations (sd),
n_alloc() distributes a total sample across strata. This is
the second half of a stratified design: strata_bound()
finds the cut points, and n_alloc() allocates within
strata.
frame <- data.frame(
N = c(4000, 3000, 3000),
sd = c(10, 15, 8),
mean = c(50, 60, 55)
)
# Fixed total n with Neyman allocation
n_alloc(frame, n = 600, alloc = "neyman")
#> Stratum allocation (neyman, 3 strata)
#> n = 600, cv = 0.0079, se = 0.4305Three solve modes are available: fixed total n, target
cv, or budget constraint (when
cost is provided in the frame).
Constraints and domain-level targets are often the more practical use cases:
frame_constraints <- transform(
frame,
cost = c(1, 1.5, 1),
max_weight = c(25, 20, NA),
take_all = c(FALSE, FALSE, TRUE)
)
n_alloc(frame_constraints, budget = 3500, alloc = "optimal", min_n = 40)
#> Stratum allocation (optimal, 3 strata)
#> n = 3404, cv = 0.0076, se = 0.4125
#> (min_n = 40)
frame_domains <- data.frame(
province = c("North", "North", "South", "South"),
stratum = c("Urban", "Rural", "Urban", "Rural"),
N = c(2000, 3000, 1800, 3200),
sd = c(12, 18, 10, 16),
mean = c(55, 48, 58, 50)
)
# Minimum total n such that each province meets the CV target
n_alloc(frame_domains, domains = "province",
cv = 0.04, alloc = "power", power_q = 0.3)
#> Stratum allocation (power, 4 strata)
#> n = 111, cv = 0.0272, se = 1.4076
#> Domains: 2
#> ---
#> province .domain .n .se .moe .cv .cost
#> North North 59.23404 2.032000 3.982647 0.0400 59
#> South South 51.50815 1.948447 3.818886 0.0368 52prec_alloc() computes the achieved precision for any
allocation:
prec_alloc(frame, n = c(200, 250, 150))
#> Sampling precision for alloc
#> n = 200 n = 250 n = 150
#> se = 0.4321, moe = 0.8469, cv = 0.0079Power analysis
Precision analysis answers “how precisely can we estimate a level?” Power analysis answers a different question: “can we detect a change or difference between two groups or time points?”
Sample size for a detectable change
A survey measured vaccination coverage at 70%. How many households are needed in a follow-up to detect a 5 percentage point increase with 80% power?
power_prop(p1 = 0.70, p2 = 0.75, power = 0.80)
#> Power analysis for proportions (solved for sample size)
#> n = 1248 (per group), power = 0.800, effect = 0.0500
#> (p1 = 0.700, p2 = 0.750, alpha = 0.05)With a design effect:
power_prop(p1 = 0.70, p2 = 0.75, power = 0.80, deff = 2.0)
#> Power analysis for proportions (solved for sample size)
#> n = 2496 (per group), power = 0.800, effect = 0.0500
#> (p1 = 0.700, p2 = 0.750, alpha = 0.05, deff = 2.00)Three solve modes
power_prop() and power_mean() each solve
for whichever of n, power, or the effect size
is left unspecified:
# Solve for n (default when p2 and power given)
power_prop(p1 = 0.70, p2 = 0.75, power = 0.80, deff = 2.0)
#> Power analysis for proportions (solved for sample size)
#> n = 2496 (per group), power = 0.800, effect = 0.0500
#> (p1 = 0.700, p2 = 0.750, alpha = 0.05, deff = 2.00)
# Solve for power (set power = NULL)
power_prop(p1 = 0.70, p2 = 0.75, n = 1500, power = NULL, deff = 2.0)
#> Power analysis for proportions (solved for power)
#> n = 1500 (per group), power = 0.584, effect = 0.0500
#> (p1 = 0.700, p2 = 0.750, alpha = 0.05, deff = 2.00)
# Solve for MDE (omit p2)
power_prop(p1 = 0.70, n = 1500, deff = 2.0)
#> Power analysis for proportions (solved for minimum detectable effect)
#> n = 1500 (per group), power = 0.800, effect = 0.0639
#> (p1 = 0.700, p2 = 0.764, alpha = 0.05, deff = 2.00)Panel surveys
If the follow-up resamples a fraction of the original households and the between-round correlation is known, the required sample size drops:
power_prop(p1 = 0.70, p2 = 0.75, power = 0.80, deff = 2.0,
overlap = 0.50, rho = 0.6)
#> Power analysis for proportions (solved for sample size)
#> n = 1749 (per group), power = 0.800, effect = 0.0500
#> (p1 = 0.700, p2 = 0.750, alpha = 0.05, deff = 2.00, overlap = 0.50, rho = 0.60)Continuous outcomes
The same three solve modes work for means:
# Sample size to detect a difference of 5 (within-group var = 200)
power_mean(effect = 5, var = 200)
#> Power analysis for means (solved for sample size)
#> n = 126 (per group), power = 0.800, effect = 5.0000
#> (alpha = 0.05)
# MDE with n = 400 per group
power_mean(var = 200, n = 400)
#> Power analysis for means (solved for minimum detectable effect)
#> n = 400 (per group), power = 0.800, effect = 2.8016
#> (alpha = 0.05)Difference-in-differences
power_did() handles two-group, two-period DiD designs.
Specify treated and control group outcomes as
c(baseline, endline) vectors:
# Proportion outcome: treated improves, control stable
power_did(treat = c(0.50, 0.55), control = c(0.50, 0.48),
outcome = "prop", effect = 0.07)
#> Power analysis for DiD proportions (solved for sample size)
#> n = 1598 (per group), power = 0.800, effect = 0.0700
#> (treat = (0.500, 0.550), control = (0.500, 0.480), alpha = 0.05)
# Mean outcome with panel overlap
power_did(treat = c(50, 55), control = c(50, 52),
outcome = "mean", var = 100, effect = 3,
overlap = 0.5, rho = 0.6)
#> Power analysis for DiD means (solved for sample size)
#> n = 245 (per group), power = 0.800, effect = 3.0000
#> (treat = (50.000, 55.000), control = (50.000, 52.000), alpha = 0.05, var = (100.00, 100.00, 100.00, 100.00), overlap = 0.50, rho = 0.60)Unequal groups and alternative methods
Both power_prop() and power_mean() support
unequal group sizes via ratio or explicit
n = c(n1, n2):
# 2:1 allocation ratio
power_prop(p1 = 0.30, p2 = 0.35, ratio = 2)
#> Power analysis for proportions (solved for sample size)
#> n1 = 2088, n2 = 1044 (total = 3132), power = 0.800, effect = 0.0500
#> (p1 = 0.300, p2 = 0.350, alpha = 0.05, ratio = 2)
# Unequal group variances
power_mean(effect = 5, var = c(80, 120))
#> Power analysis for means (solved for sample size)
#> n = 63 (per group), power = 0.800, effect = 5.0000
#> (alpha = 0.05)For rare proportions, arcsine and log-odds transforms are more accurate:
power_prop(p1 = 0.15, p2 = 0.18, alternative = "one.sided",
method = "arcsine")
#> Power analysis for proportions (solved for sample size)
#> n = 1890 (per group), power = 0.800, effect = 0.0300
#> (p1 = 0.150, p2 = 0.180, alpha = 0.05, one-sided, method = arcsine)
power_prop(p1 = 0.15, p2 = 0.18, alternative = "one.sided",
method = "logodds")
#> Power analysis for proportions (solved for sample size)
#> n = 1889 (per group), power = 0.800, effect = 0.0300
#> (p1 = 0.150, p2 = 0.180, alpha = 0.05, one-sided, method = logodds)Power curve
plot() draws the power-vs-sample-size curve with
reference lines at the solved point:
pw <- power_prop(p1 = 0.70, p2 = 0.75, power = 0.80, deff = 2.0)
plot(pw)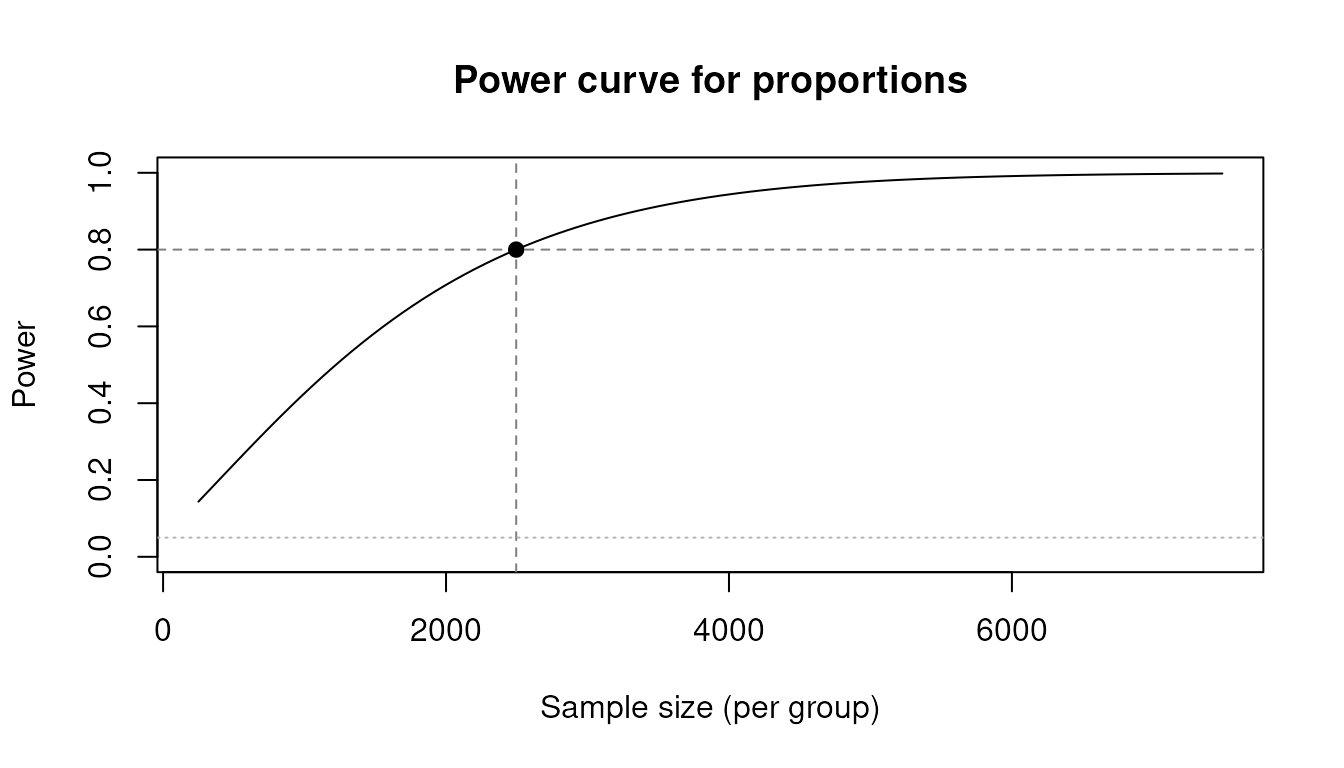
Sensitivity analysis
How sensitive is a sample size to assumptions about the design
effect, response rate, or prevalence? The predict() method
evaluates any svyplan result at new parameter
combinations.
Varying design parameters
x <- n_prop(p = 0.3, moe = 0.05, deff = 1.5)
predict(x, expand.grid(
deff = c(1.0, 1.5, 2.0, 2.5),
resp_rate = c(0.8, 0.9, 1.0)
))
#> deff resp_rate n se moe cv
#> 1 1.0 0.8 403.3532 0.02551067 0.05 0.08503558
#> 2 1.5 0.8 605.0298 0.02551067 0.05 0.08503558
#> 3 2.0 0.8 806.7064 0.02551067 0.05 0.08503558
#> 4 2.5 0.8 1008.3829 0.02551067 0.05 0.08503558
#> 5 1.0 0.9 358.5362 0.02551067 0.05 0.08503558
#> 6 1.5 0.9 537.8042 0.02551067 0.05 0.08503558
#> 7 2.0 0.9 717.0723 0.02551067 0.05 0.08503558
#> 8 2.5 0.9 896.3404 0.02551067 0.05 0.08503558
#> 9 1.0 1.0 322.6825 0.02551067 0.05 0.08503558
#> 10 1.5 1.0 484.0238 0.02551067 0.05 0.08503558
#> 11 2.0 1.0 645.3651 0.02551067 0.05 0.08503558
#> 12 2.5 1.0 806.7064 0.02551067 0.05 0.08503558Cluster budget scenarios
cl <- n_cluster(stage_cost = c(500, 50), delta = 0.05, budget = 100000)
predict(cl, data.frame(budget = c(50000, 100000, 150000, 200000)))
#> budget n_psu psu_size total_n cv cost
#> 1 50000 42.04499 13.78405 579.5501 0.05318275 50000
#> 2 100000 84.08997 13.78405 1159.1003 0.03760588 100000
#> 3 150000 126.13496 13.78405 1738.6504 0.03070507 150000
#> 4 200000 168.17994 13.78405 2318.2006 0.02659137 200000Power curves
pw <- power_prop(p1 = 0.30, p2 = 0.35, n = 500, power = NULL)
predict(pw, data.frame(n = seq(200, 1000, 200)))
#> n power effect
#> 1 200 0.1877131 0.05
#> 2 400 0.3272968 0.05
#> 3 600 0.4569385 0.05
#> 4 800 0.5707088 0.05
#> 5 1000 0.6665884 0.05Precision by sample size
pr <- prec_prop(p = 0.3, n = 400)
predict(pr, data.frame(n = c(100, 200, 400, 800, 1600)))
#> n se moe cv
#> 1 100 0.04582576 0.08981683 0.15275252
#> 2 200 0.03240370 0.06351009 0.10801234
#> 3 400 0.02291288 0.04490842 0.07637626
#> 4 800 0.01620185 0.03175505 0.05400617
#> 5 1600 0.01145644 0.02245421 0.03818813Putting it all together
A typical planning sequence for a national household survey ties
several svyplan functions together. A svyplan() profile
captures the shared design assumptions:
- Set shared design parameters once with
svyplan() - Define indicators and precision targets per domain
- Compute sample sizes with
n_multi() - Evaluate achieved precision with
prec_multi() - Run sensitivity analysis with
predict() - If frame data is available, estimate variance components with
varcomp() - Optimize the multistage allocation with
n_cluster()orn_multi()with costs - Determine strata boundaries with
strata_bound() - Check that the design has adequate power with
power_prop()/power_mean()
# 1: shared design assumptions
design <- svyplan(deff = 2.0, resp_rate = 0.85)
# 2-3: multi-indicator sample size
targets <- data.frame(
name = c("stunting", "vaccination", "anemia"),
p = c(0.25, 0.70, 0.12),
moe = c(0.05, 0.05, 0.03),
deff = c(2.0, 1.5, 2.5)
)
result <- n_multi(targets)
result
#> Multi-indicator sample size
#> n = 1127 (binding: anemia)
#> ---
#> name .n .cv_target .cv_achieved .binding
#> stunting 577 0.10204269 0.07297042
#> vaccination 485 0.03644382 0.02388518
#> anemia 1127 0.12755336 0.12755336 *
# 4: achieved precision for each indicator
prec_multi(result)
#> Multi-indicator sampling precision
#> name .se .moe .cv
#> stunting 0.02551067 0.05 0.10204269
#> vaccination 0.02551067 0.05 0.03644382
#> anemia 0.01530640 0.03 0.12755336
# 5: how sensitive is the binding indicator to the response rate?
predict(
n_prop(p = 0.25, moe = 0.05, plan = design),
data.frame(resp_rate = c(0.7, 0.8, 0.9, 1.0))
)
#> resp_rate n se moe cv
#> 1 0.7 823.1697 0.02551067 0.05 0.1020427
#> 2 0.8 720.2735 0.02551067 0.05 0.1020427
#> 3 0.9 640.2431 0.02551067 0.05 0.1020427
#> 4 1.0 576.2188 0.02551067 0.05 0.1020427
# 9: can we detect a 5pp decline in stunting with this sample?
power_prop(
p1 = 0.25, p2 = 0.20,
n = as.integer(result), power = NULL,
plan = design
)
#> Power analysis for proportions (solved for power)
#> n = 1127 (net: 958, per group), power = 0.459, effect = 0.0500
#> (p1 = 0.250, p2 = 0.200, alpha = 0.05, deff = 2.00, resp_rate = 0.85)All svyplan output objects support
as.integer() (ceiling of n) and
as.double() (exact n), making it straightforward
to pass results between functions or into other packages.